De Novo Genome Assembly (De Novo Genome Assembly)
Jump to navigation
Jump to search
Description
De novo sequencing refers to sequencing a novel genome where there is no reference sequence available for alignment. Sequence reads are assembled as contigs, and the coverage quality of de novo sequence data depends on the size and continuity of the contigs (ie, the number of gaps in the data).
Parameters
$n$: sum of lengths of reads
$f$: number of input sequences
Table of Algorithms
| Name | Year | Time | Space | Approximation Factor | Model | Reference |
|---|---|---|---|---|---|---|
| Overlap Layout Consensus | 1987 | $O(n^{2})$ | $O(n^{2})$? | Deterministic | ||
| Greedy SEQAID | 1984 | $O(n^{2})$? | $O(n^{2})$? | Deterministic | Time | |
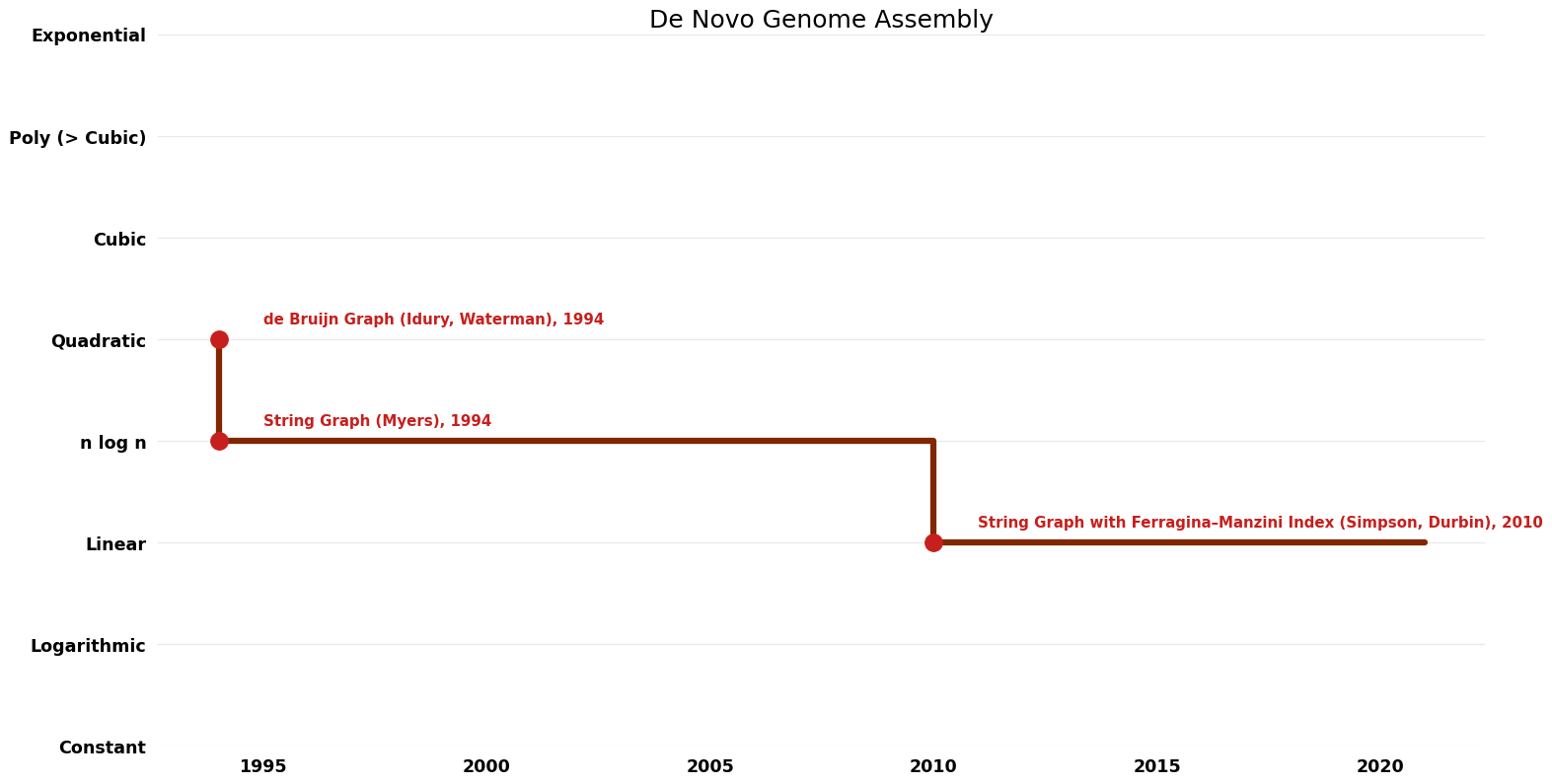
| de Bruijn Graph (Idury, Waterman) | 1994 | $O(n^{2})$ | $O(n)$? | Exact | Deterministic | Time |
| String Graph (Myers) | 1994 | $O(n \log n)$ | $O(n)$? | Exact | Deterministic | Time |
| String Graph with Ferragina–Manzini Index (Simpson, Durbin) | 2010 | $O(n)$ | $O(n)$? | Exact | Deterministic | Time |
| Hybrid Algorithm | 1999 | $O(n^{2})$ | Exact | Deterministic |